Agents for biology are still stuck in the demo-to-deployment gap. With SpatialBench, we showed that frontier models struggle to extract biological insight from spatial transcriptomics data. But spatial is only part of the story. Single-cell RNA sequencing is the workhorse assay of modern biology, far more widely adopted, with more public data, more mature tooling, and more documentation for models to train on. If agents can’t handle scRNA-seq, the path to reliable agentic data analysis is far away.
Here we introduce scBench (read the paper), a benchmark of 394 verifiable problems drawn from real scRNA-seq workflows. It spans six sequencing platforms, seven task categories, and eight frontier models. The best model reaches 52.8% accuracy. Better than spatial, but it means the best agent in the world still fails nearly half the time on routine analysis tasks.
Thanks for reading! Subscribe for free to receive new posts and support my work.
scBench tests what bioinformaticians actually do
Existing biology benchmarks reward textbook recall or literature-style reasoning. Real analysis requires loading messy datasets, writing code, making judgment calls about thresholds and parameters, and producing concrete quantitative results. scBench recreates these conditions.
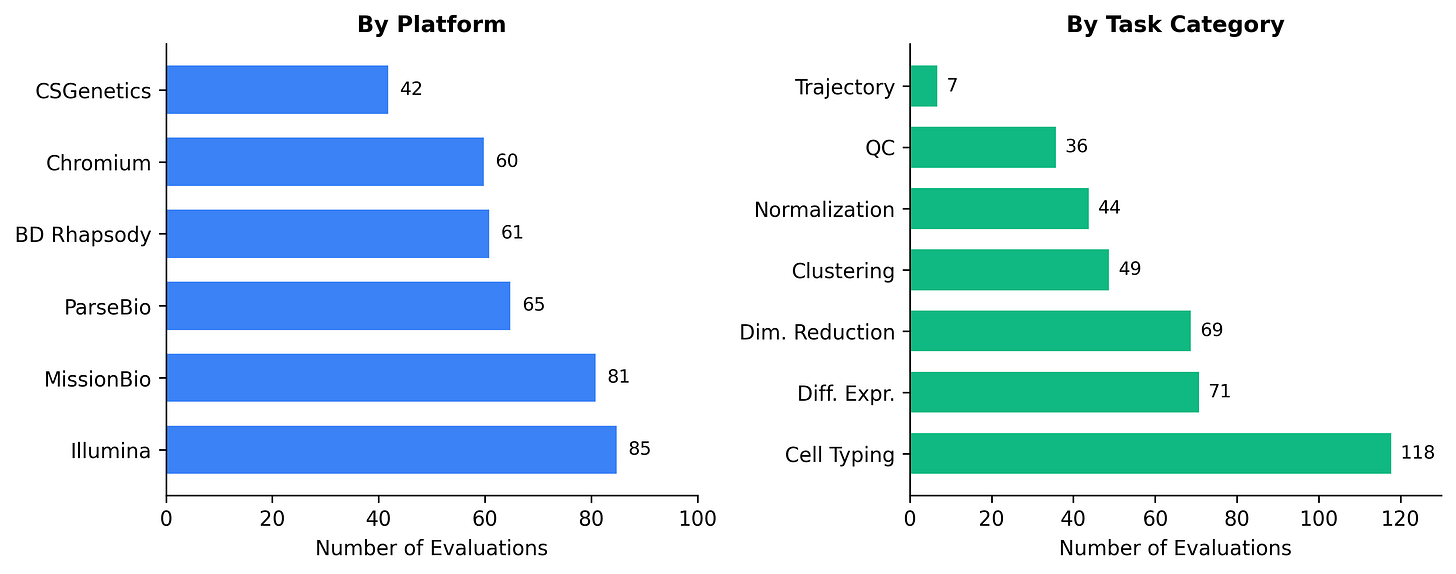
The benchmark covers 394 problems across six sequencing platforms (Chromium, BD Rhapsody, CSGenetics, Illumina, MissionBio, ParseBio) and seven task categories (QC, normalization, dimensionality reduction, clustering, cell typing, differential expression, and trajectory analysis). Each problem provides a data snapshot – typically an AnnData `.h5ad` file-captured right before a decision point, a natural-language task prompt, and a deterministic grader that scores the agent’s structured JSON output as pass or fail.
We pressure-tested each problem for shortcuts: removing precomputed embeddings, stripping cached labels, and ensuring answers can’t be guessed from prior knowledge alone. Cell typing (118 problems, 30%) and differential expression (71, 18%) dominate the benchmark because these are the stages where dataset-specific judgment matters most and where agents struggle hardest.
Agents are better at scRNA-seq than spatial – but still not reliable
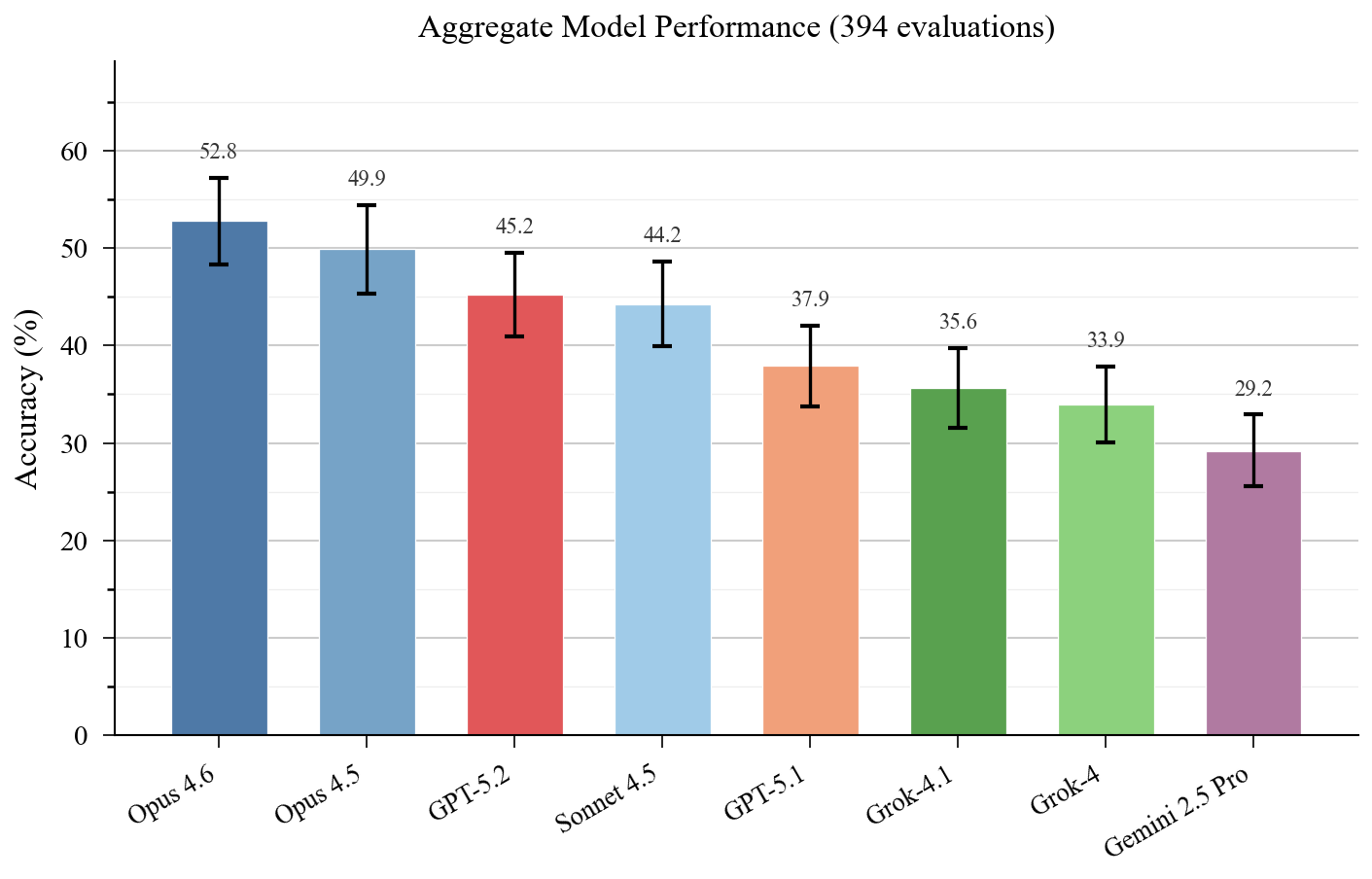
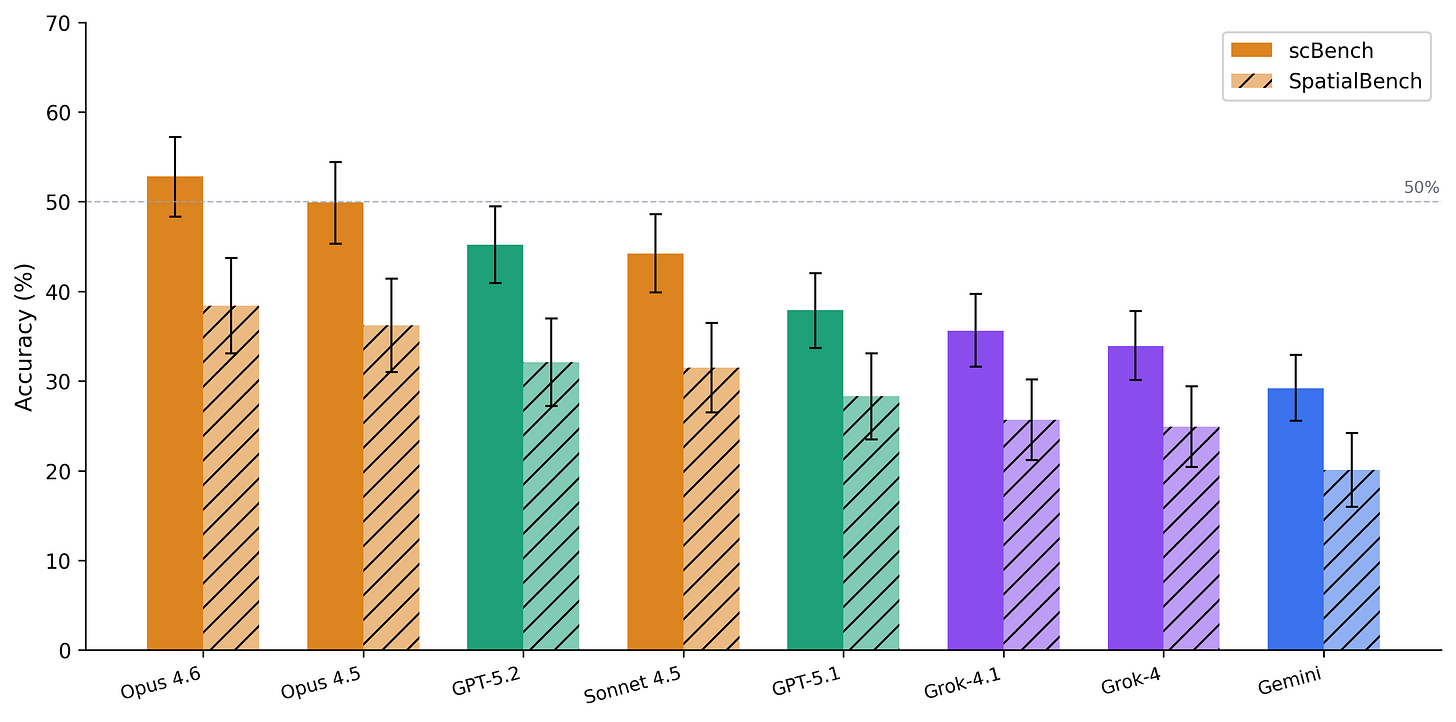
Across eight frontier models from four providers, accuracy ranges from 29% to 53%.
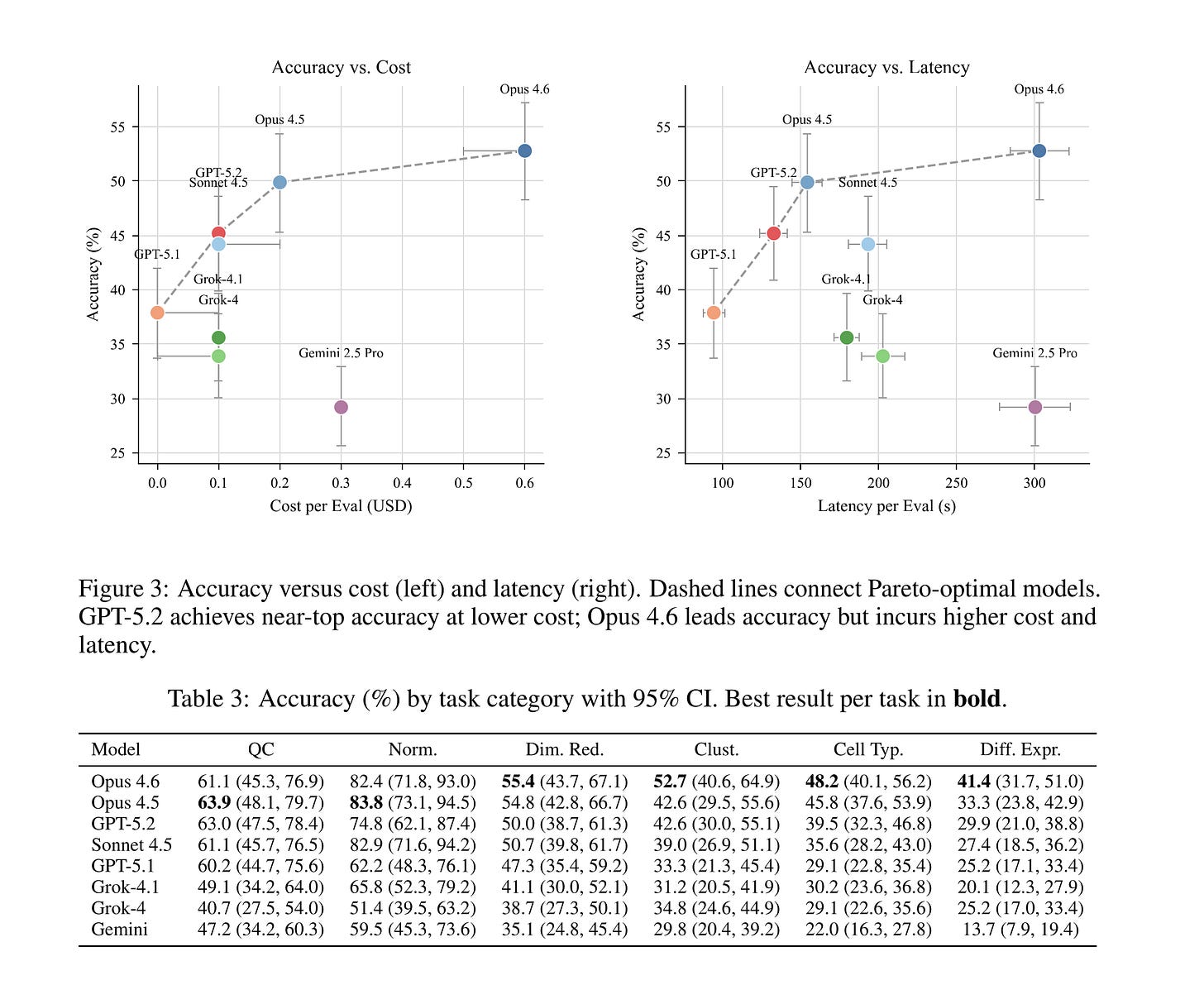
Claude Opus 4.6 leads at 52.8%, followed by Opus 4.5 (49.9%), GPT-5.2 (45.2%), and Sonnet 4.5 (44.2%). The bottom tier – GPT-5.1 (37.9%), Grok-4.1 (35.6%), Grok-4 (33.9%), and Gemini 2.5 Pro (29.2%) – trails by a wide margin. The 23.6 percentage point spread between best and worst exceeds SpatialBench’s 18.3 point spread, meaning scBench discriminates model capability even at higher overall accuracy.
The accuracy bump over SpatialBench likely reflects training data. scRNA-seq has far more public datasets and Scanpy dominates the ecosystem with extensive documentation and tutorials.
Tasks reveal a consistent difficulty gradient
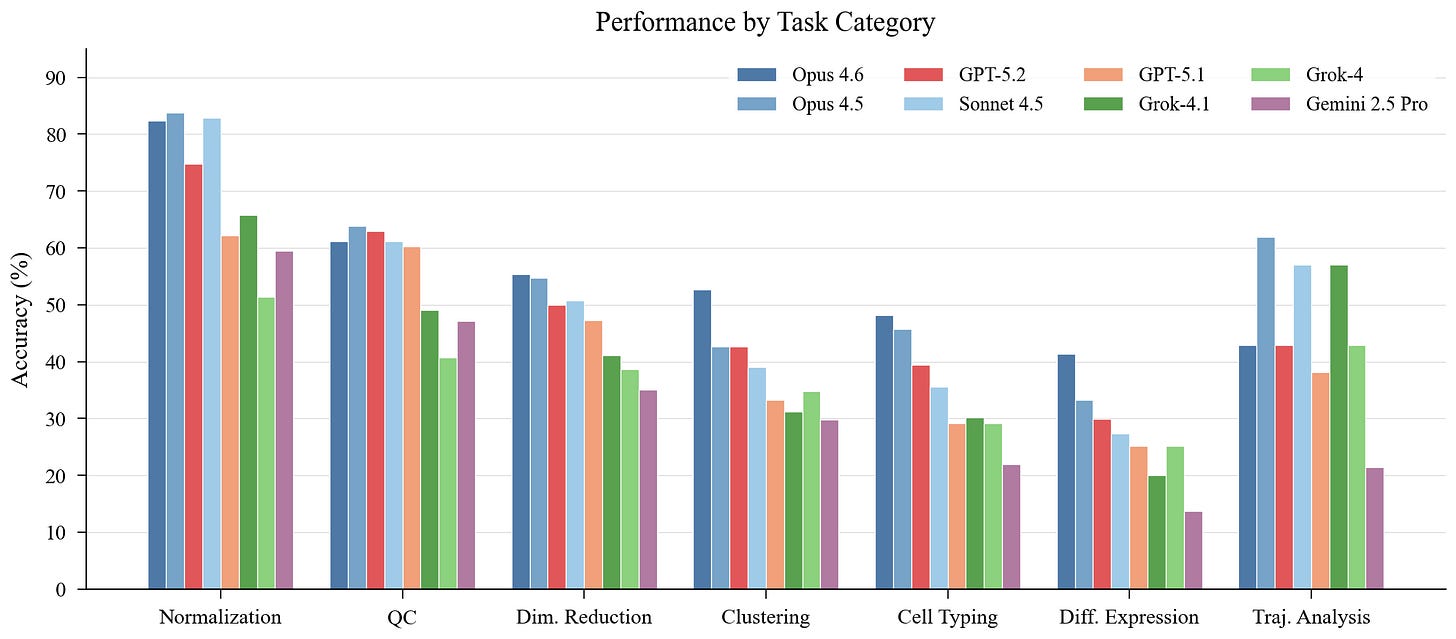
Not all analysis steps are created equal. Normalization is easiest (cross-model mean 70.4%), followed by QC (55.3%). These are procedural—apply a well-known transformation, check a metric against a threshold. Agents handle them reasonably well.
The story changes for judgment-heavy tasks. Clustering drops to 38.3%, cell typing to 34.9%, and differential expression to 27.0%. Seven of eight models follow this same difficulty ordering. DE is also where models diverge most: 27.7 percentage points separate best from worst.
The pattern is clear. Tasks that require contextual scientific reasoning-selecting marker genes, interpreting cluster identity, choosing statistical tests, identifying tissue-specific signatures-are where agents break down. General-purpose coding skill is necessary but not sufficient.
Platform choice matters as much as model choice
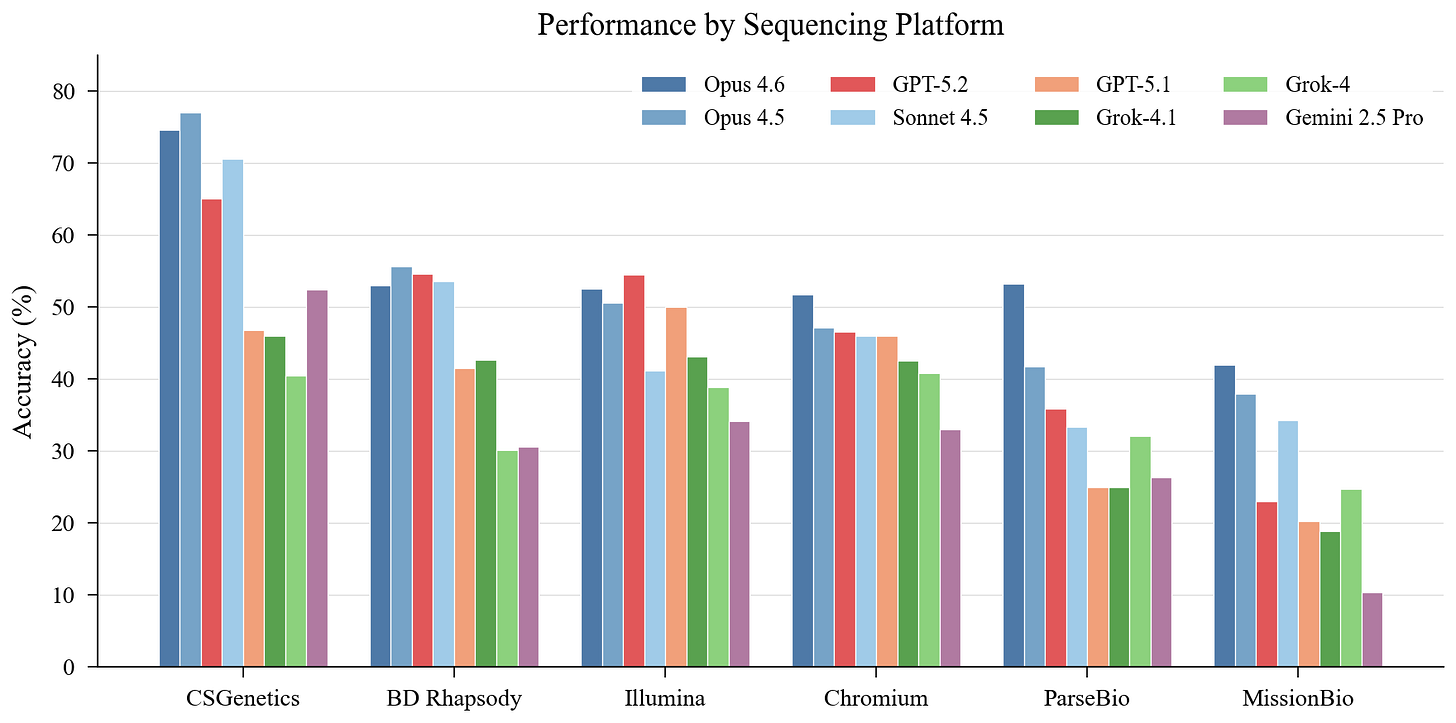
This is perhaps the most striking finding. Cross-model mean accuracy ranges from 59.1% on CSGenetics to 26.4% on MissionBio—a 32.7 point gap that exceeds the 23.6 point spread between best and worst models. Platform matters more than which model you use.
CSGenetics is easiest for six of eight models. MissionBio is hardest for all eight. The MissionBio collapse is dramatic: even the best model (Opus 4.6) only hits 42%, and Gemini drops to 10.3%. Every model shows large platform swings—Gemini drops 42 points between its best and worst platform. Even Opus 4.5, the most consistent model, loses 39 points.
MissionBio inverts the overall rankings in interesting ways. Grok-4 (sixth overall) beats GPT-5.2 (third overall) on MissionBio. Sonnet 4.5 surpasses GPT-5.2 by 11 points. Models that memorized Scanpy tutorials without learning transferable analysis techniques collapse on platforms with non-standard data structures and less common tooling.
These effects almost certainly reflect training data composition. Chromium and Illumina dominate public repositories and documentation. MissionBio and ParseBio appear less frequently. Reliable agents will need platform-aware context and assay-specific tooling, not one-size-fits-all reasoning.
scBench complements SpatialBench across single-cell modalities
Together, scBench and SpatialBench cover the two dominant transcriptional assays. The top model reaches 52.8% on scBench versus 38.4% on SpatialBench-scRNA-seq is more tractable. But the structural patterns are shared: normalization is easiest in both, platform effects drive 30–40 point swings in both, and model rankings are preserved at the extremes (Claude Opus leads both, Gemini ranks last in both).
The benchmarks form a complementary diagnostic. scBench tests whether models can handle the most common and best-documented assay type. SpatialBench tests whether that competence transfers to newer, less standardized technologies. Together they reveal whether an agent has learned general analysis reasoning or just memorized Scanpy workflows.
Limitations
Deterministic grading enables verifiable evaluation but necessarily discretizes scientific judgment into automatically checkable chunks. Each evaluation snapshots a single workflow step rather than capturing the long-horizon iteration where errors compound and thresholds are revisited. Real analysis is messier, more iterative, and longer than any single problem captures. But measuring step-level competence reliably is a prerequisite for automating longer workflows.
Where this is going
scBench confirms the pattern SpatialBench established: agents for biology are in a capability regime where they can accelerate routine analysis but cannot yet be trusted to autonomously answer scientific questions without human oversight. The path forward is a long tail of tractable engineering: platform-specific context, better harness design, and exposure to representative workflows across diverse biological contexts.
We’re building toward a family of benchmarks spanning major biological modalities, each an evolving formalization of the tacit knowledge and judgment that working scientists bring to data analysis. The goal is test-driven development of agent systems that improve through both model training and harness engineering.
Code, canonical evaluations, and full trajectories are available at github.com/latchbio/scbench.
Thanks for reading! Subscribe for free to receive new posts and support my work.